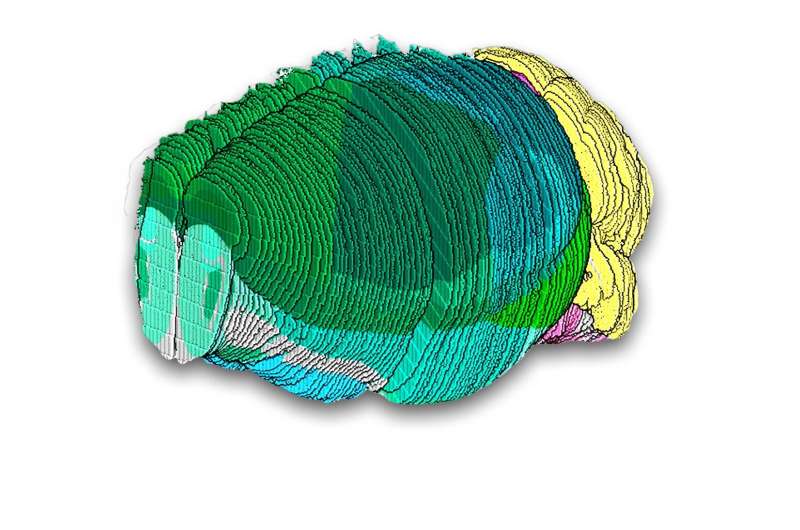
A team of scientists at the Broad Institute of MIT and Harvard has generated one of the first comprehensive maps of cell types in a mammalian brain using recently developed technology called spatial transcriptomics, which can reveal not just the gene activity of individual cells, but also their location within tissues and organs.
The atlas, described in Nature, delineates several thousand cell populations across the entire mouse brain, revealing surprising cellular diversity in understudied brain regions and offering deeper views of brain structures than were possible before.
The effort was led by Evan Macosko, an institute member at the Broad and associate professor and attending psychiatrist at Massachusetts General Hospital, and Fei Chen, a core institute member at the Broad and an assistant professor in the Department of Stem Cell and Regenerative Biology at Harvard University. Both are senior authors on the paper.
The team measured the activity of all genes in the genome of individual cells throughout the mouse brain, and assigned the cells’ locations within the tissue. Their analysis uncovered an estimated 90% of all cell populations in the mouse brain.
The scientists found most cellular diversity within relatively understudied subcortical areas of the brain, especially the midbrain, pons, medulla, and the hypothalamus. “We suspected the most diversity would be found in these areas, so we prioritized them in our profiling,” said Macosko.
“A lot of the real nuts and bolts stuff that a brain is doing is in these basic areas, which have received very little attention compared to the cortex. Our results underscore the need to study them more deeply.” The researchers also discovered clues about cellular function and the potential roles of brain structures in disease.
“Efforts like these generate crucial resources for the neuroscience community, because the brain is so enormously complicated,” said Chen.
Macosko added, “Our atlas represents the culmination of a decade of work at the Broad. Fei and I developed the technology in our labs and used it to process the largest single-cell and spatial dataset ever generated, leading to the first comprehensive atlas of cell types in a mammalian brain.”
The study is one of a package of 10 papers in Nature that take distinct, yet complementary, approaches to mapping the mouse nervous system at the single-cell level. The studies are from groups at the Broad, Allen Institute for Brain Science, the Salk Institute for Biological Studies, and other institutions that are part of the National Institutes of Health’s Brain Research Through Advancing Innovative Neurotechnologies Initiative, or The BRAIN Initiative—Cell Census Network (BICCN). Together the papers describe the first complete cell type atlas of a whole mammalian brain.
The package includes a spatial, single-cell atlas of the mouse brain and spinal cord that was first published online in September in Nature and was led by Xiao Wang, a Broad core institute member, a Merkin Institute Fellow, and an assistant professor of chemistry at MIT, and Jia Liu, an assistant professor of engineering at Harvard University.
Independently of BICCN, Wang’s and Liu’s team used a spatial transcriptomics technology they developed to analyze the expression of more than 1,000 genes and detail the locations of individual cells with unparalleled sub-cellular resolution.
They identified hundreds of cell types, generated highly precise tissue maps of the mouse brain and spinal cord, and demonstrated their method’s utility in revealing which cell types and brain regions are altered by gene-delivery viruses.
Mapping the brain
In the study from the Macosko and Chen labs, the scientists worked together to develop their spatial transcriptomic approach and apply it across the entire mouse brain. They also relied on the expertise of members of the Broad’s Genomics Platform and the team that supports Hail, an open-source tool for scalable genomic analysis.
The researchers identified several thousand unique cell populations, and mapped their locations in the whole tissue with near-cellular resolution. They estimated that their analysis captured about 90 percent of cell types in the mouse brain, including a large diversity in underexplored areas of the brain. The researchers created an online browser to house and share their datasets with the scientific community (http://www.BrainCellData.org/).
The team analyzed the full transcriptome of cells from nearly 100 regions across the mouse brain using high-throughput single-nucleus RNA sequencing, the preferred approach for efforts to create a human brain atlas. This resulted in more than four million profiles of gene activity, which they clustered into nearly 5,000 unique cell populations, most of which were neuronal cells.
The team next applied an approach developed in the Chen and Macosko labs known as Slide-seq. They transferred 101 serial sections, spanning the volume of a single mouse brain, onto arrays of beads covered in unique DNA barcodes, which bound to the mRNA transcripts in the brain tissue.
They then sequenced those transcripts and aligned that spatial data to an existing 3D reference atlas, enabling them to assign each transcript to a known brain structure, representing more than 1.7 million mapped cells.
The researchers combined detailed and well-sampled cell type profiles from the single-nucleus sequencing dataset to locate each cell type in each slice to generate a detailed and thorough atlas of the entire mouse brain. They also analyzed the dataset to come up with two or three marker genes that could be used to uniquely identify almost all of the cell populations.
The authors said that other scientific groups can use the transcriptomic profiles, spatial localizations, and sets of marker genes they identified to study particular cells of interest. Groups may also use the data by integrating it with their own more detailed examination of a particular brain region.
“We hope that our atlas can both empower the community in their own work and allow them to further explore the cell types we identified,” said co-first author Jonah Langlieb, a computational biologist in the Macosko lab who led the work along with co-first author Nina Sachdev.
The team also revealed how signaling molecules known as neurotransmitters are used by different cell types in various brain regions. In addition, they demonstrated their atlas’s utility in revealing where disease-associated genes are active, for example, specific neuronal cells that are enriched for expression of genetic factors associated with schizophrenia.
Together, the 10 studies provide rich and profound new views of the remarkable complexity of the brain, and their datasets and methods set the stage for larger efforts to do the same for other mouse organs and for the human brain.
“By mapping the mouse brain, the primary mammalian system used in neuroscience, we’ve provided molecular, functional, and anatomical classifications that provide a key foundation for human brain mapping, which is what comes next,” said Macosko.
More information:
Hailing Shi et al, Spatial atlas of the mouse central nervous system at molecular resolution, Nature (2023). DOI: 10.1038/s41586-023-06569-5
Langlieb J, Sachdev NS, et al, The cell type composition of the adult mouse brain revealed by single cell and spatial genomics, Nature (2023). On bioRxiv: DOI: 10.1101/2023.03.06.531307
Source: Read Full Article