In a recent study published in the Nature Communications journal, researchers monitored the changing symptom profiles associated with different severe acute respiratory syndrome CoV-2 (SARS-CoV-2) variants, especially Omicron subvariants BA.1 and BA.2.
 Study: Variant-specific symptoms of COVID-19 in a study of 1,542,510 adults in England. Image Credit: NIAID
Study: Variant-specific symptoms of COVID-19 in a study of 1,542,510 adults in England. Image Credit: NIAID
Background
Previous community-based studies have shown that symptom profiles differ for different SARS-CoV-2 variants. So they used variable selection and ranking techniques to identify symptoms that were predictive of the causal variant. They also assessed how much symptom data could predict SARS-CoV-2 reverse transcription-polymerase chain reaction (RT-PCR) positivity.
Identifying individuals more likely to be infected and infectious based on symptom profiles have much clinical value. With many countries having already lifted restrictions, mandatory isolation measures, and routine testing, such monitoring efforts will become increasingly important.
The REal-time Assessment of Community Transmission−1 (REACT-1) study monitored the spread and clinical manifestation of SARS-CoV-2 monthly in the population in England between 1 May 2020 and 31 March 2022.
About the study
In the present study, researchers used regression and variable selection models to examine the REACT-1 study data of over 1.5 million randomly selected people from the population of England. Next, they predicted symptom profiles of each SARS-CoV-2 variant that was dominant in England and worldwide over this period. These variants were wild-type (wt) ancestral strain, Alpha, Delta, and Omicron BA.1 and BA.2. More importantly, for each variant, they identified 26 symptoms most predictive of high viral load, indicating higher infectiousness and ability to more quickly transmit COVID-19.
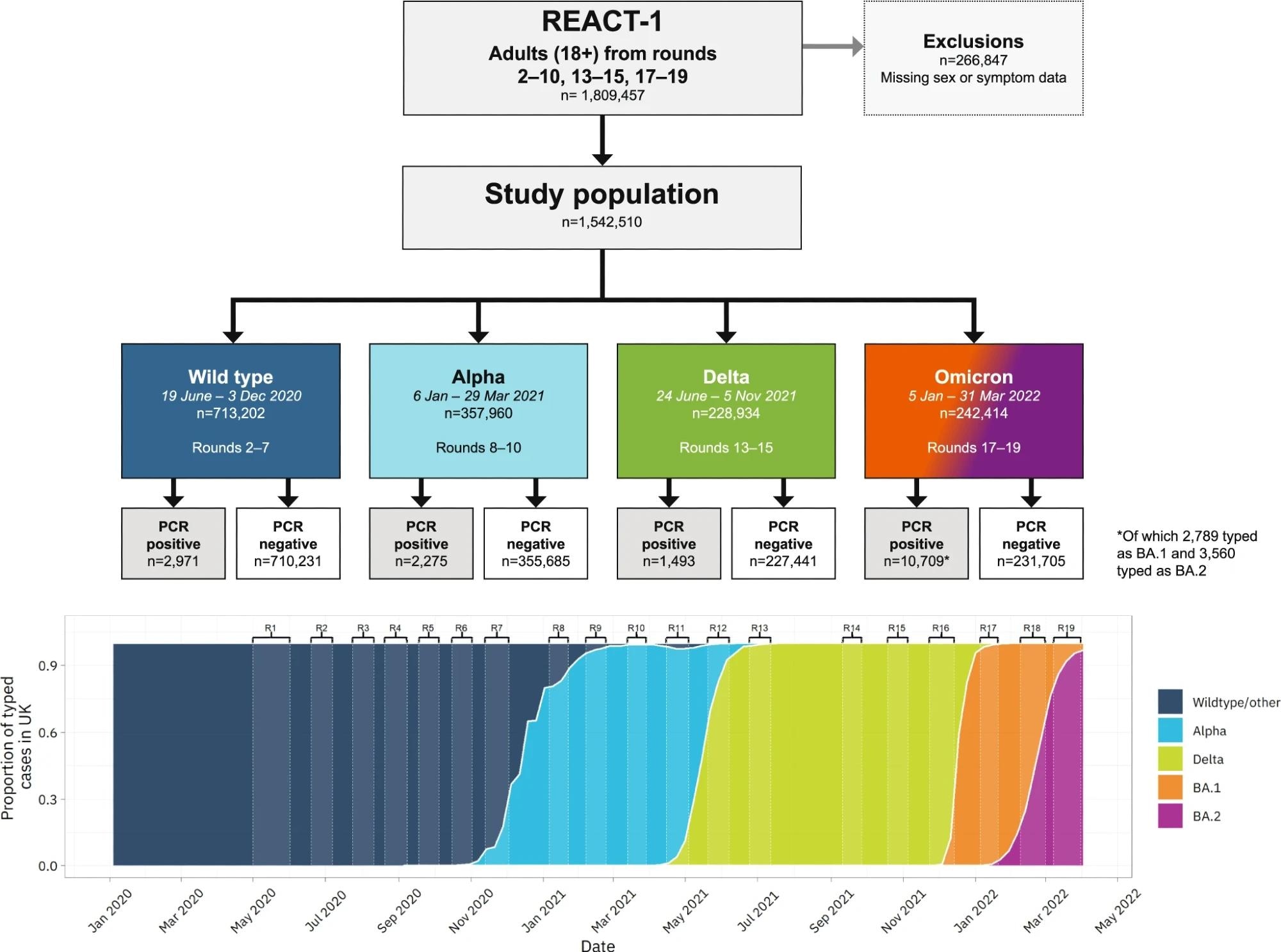 Variant prevalence data in the bottom panel is from GISAID.
Variant prevalence data in the bottom panel is from GISAID.
The study population comprised 1,542,510 adults aged 18 or above, including 17,448 individuals positive for SARS-CoV-2. Of these, 2971, 2275, 1493, and 10,709 individuals tested positive for wt, Alpha, Delta, and Omicron variants, respectively. In multivariable analysis, the team used Least Absolute Shrinkage and Selection Operator (LASSO) penalized logistic regression for identifying all symptoms positively predictive of COVID-19 positivity for each variant. This method accounted for the differences in symptoms by variant type. In addition, it considered cold-like symptoms, such as running nose and sneezing, for Omicron BA.1 and BA.2.
Study findings
The researchers observed differing underlying pathophysiology associated with different SARS-CoV-2 variants. Against the background of differing infection- and vaccine-induced population immunity, there were differences in coronavirus disease 2019 (COVID-19) symptoms due to Omicron compared with previous variants and within Omicron (BA.2 vs. BA.1). Individuals infected with Omicron BA.2 reported the most of any of the 26 symptoms (75.9%), followed by BA.1 (70%), Delta (63.8%), Alpha (54.7%), and wt (45%).
Loss of sense of smell and taste and cold-like symptoms were less and more predictive of swab positivity for Omicron than for other variants, respectively. Specifically, Omicron infections were not so strongly associated with anosmia as previous variants. Likely, alterations in the viral gene sequences that regulate host responses in individuals infected with Omicron reduced the downregulation of expression of olfactory receptors, which caused anosmia following COVID-19. However, comprehensive transcriptomic studies in animals and humans could identify and elucidate the mechanisms involved in this phenomenon.
Furthermore, the researchers noted that while Omicron BA.2 infections were more likely to be symptomatic, with cold- or influenza-like symptoms, 54% greater odds of symptoms affecting day-to-day activities 'a lot' and on average one additional symptom, compared to BA.1. Indeed, the higher symptom burden and severity associated with BA.2 might also have a higher societal and economic impact.
Conclusions
The inherent severity of SARS-CoV-2 variants is multifaceted, owing to varying levels of population immunity due to prior SARS-CoV-2 infection or vaccination. Nevertheless, the swift replacement of BA.1 by BA.2 in England, and a high PCR positivity, allowed a comparison of the symptom burden and symptom severity of these two variants within a population with similar characteristics.
As a result of adjusting for vaccine booster status and the time since the last vaccine dose, the symptom burden and severity-related findings for Omicron BA.2 with BA.1 were robust. They also support the prior findings that Omicron BA.2 has higher transmissibility in a highly vaccinated population.
Finally, the study results revealed that Omicron infections caused symptoms such as fever, chills, sore throat, muscle aches, runny nose, sneezing, and headaches that were associated with the lowest adjusted cycle thresholds (CT). It further supports that Omicron exhibited higher viral loads and infectivity than previous variants.
Overall, based on symptom profiles reported during two years of the COVID-19 epidemic in England, Omicron subvariant BA.2 caused more symptoms and disruption to the daily activities of infected individuals than its predecessors. As a result, from 1 April 2022, the England government moved to the policy of 'living with COVID-19'. While many countries, including England, have stopped or reduced free or routine SARS-CoV-2 testing programs, new variants of the virus are still emerging. Understanding the symptom profiles may help identify high-risk individuals.
- Variant-specific symptoms of COVID-19 in a study of 1,542,510 adults in England, Matthew Whitaker, Joshua Elliott, Barbara Bodinier, Wendy Barclay, Helen Ward, Graham Cooke, Christl A. Donnelly, Marc Chadeau-Hyam, Paul Elliott, Nature Communications 2022, DOI: https://doi.org/10.1038/s41467-022-34244-2, https://www.nature.com/articles/s41467-022-34244-2#Sec7
Posted in: Child Health News | Men's Health News | Medical Science News | Medical Research News | Women's Health News | Disease/Infection News
Tags: Anosmia, Cold, Coronavirus, Coronavirus Disease COVID-19, covid-19, CT, Fever, Gene, immunity, Influenza, Muscle, Omicron, Pathophysiology, Polymerase, Polymerase Chain Reaction, Respiratory, Running, SARS, SARS-CoV-2, Severe Acute Respiratory, Severe Acute Respiratory Syndrome, Sneezing, Sore Throat, Syndrome, Throat, Transcription, Vaccine, Virus

Written by
Neha Mathur
Neha is a digital marketing professional based in Gurugram, India. She has a Master’s degree from the University of Rajasthan with a specialization in Biotechnology in 2008. She has experience in pre-clinical research as part of her research project in The Department of Toxicology at the prestigious Central Drug Research Institute (CDRI), Lucknow, India. She also holds a certification in C++ programming.
Source: Read Full Article